Big genomes
Reading: Biggest genome ever found belongs to this odd little plant, Nature, 31 May 2024.
Reading this I learned that our genome is made up of 3.2 Gbp (giga base pairs, so 3.2 billion base pairs). That’s for one set of chromosomes, “1C” counts.
For comparison, the record holder is now a New Caledonian fork fern with 160 Gbp. The largest animal is a lung fish with just under 130 Gbp.
These aren’t sequenced genomes. They are estimates from comparisons to known genome length. From the paper: “Briefly, about 1 cm of fresh leaf material of the study species and the calibration standard species were chopped together [ …] Genome sizes were estimated by calculating the ratio between the mean fluorescence of […] the calibration standard species for which the genome size is known, and the mean fluorescence […] for the nuclei from the target species”.
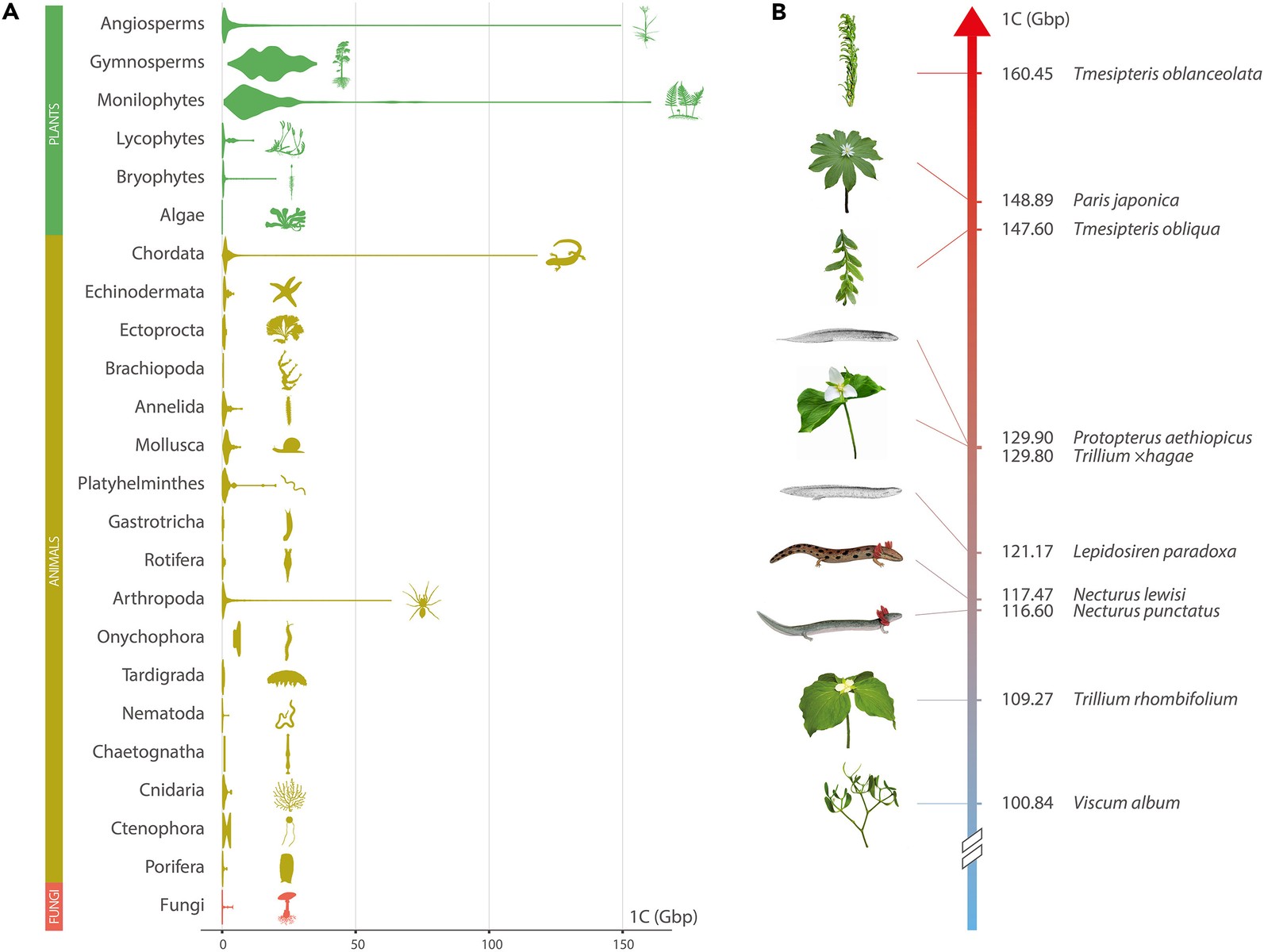
There are lots of “why?” questions to be asked of this size, and apparently:
“What controls plant genome size and complexity?” was among the top “One hundred important questions facing plant science.”
It might turn out just be neutral (not costing the plant or animal much).
I was also reading about the largest animal genome sequenced, which is a 91 Gbp lung fish. It has 20,000 genes (like us), and a load of non-coding DNA:
The large size of the lungfish genome is due precisely to the enormous abundance of repetitive elements in the genome, especially retrotransposons and transposons.
I’d love to understand the adaptive advantage of all of this. Fascinating area.
Related:
- The Animal Genome Size Database — a bit wonky in places in my browser, but “catalogue of animal genome size data”, measured in “picograms” but easy to convert to base pairs.
- Plant DNA C-values Database — from Kew.
- Fungal Genome Size Database — which also contains links to yet more genome databases.
- Darwin Tree of Life — aiming to “sequence the genomes of 70,000 species of eukaryotic organisms in Britain and Ireland”.
