I'm working on a machine learning cancer classification problem, but we have only a handful of positive cases in the data. Thankfully. But that fact does make my job harder.
It causes three main problems:
- We may not have enough variety to be able to find the true patterns in the data. This is a fundamental blocker, and the solution is: get more data, or don't use machine learning.
- When we evaluate a model, we have to trade-off what we use in training (as much as possible, please) against how much we hold back for fair evaluation (a representative selection). Cross-validation is how you tend to do this, building multiple models on different cuts of the data, to get a realistic estimate of performance. I have just about enough data to do this.
- However, the killer problem is how you find the hyperparameters for the model in the first place. Many models learn from data by adjusting parameters, but the models themselves have "settings" to control how they behave, and these are called hyperparameters (or meta-parameters). What you'd like to do is have a big set of data to train on, and as you do that, you test against a held-out data set to measure how you're doing, and to make sure you're not over-fitting (memorising) the positive cases. I don't have the data to do this.
The best practice here is stratified nested cross-validation. It's a bit involved, as I'll illustrate.
Here's the sketch I made myself for how this works:
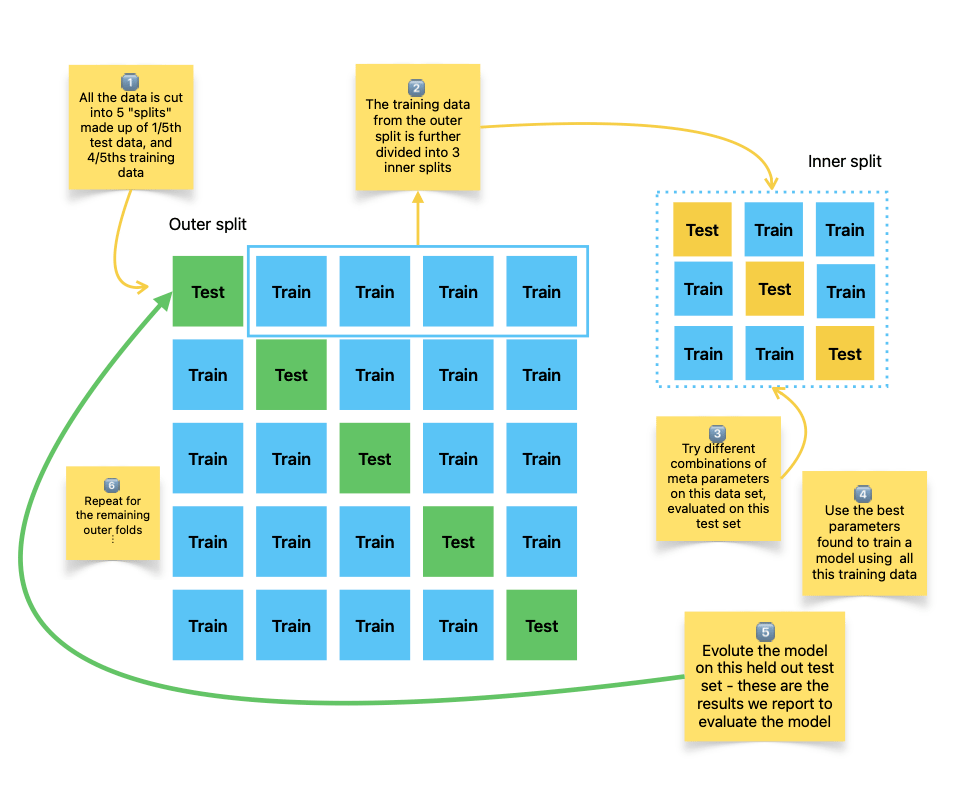
The big main square of green and blue boxes is how standard k-fold cross-validation looks. We divide the data into five (k=5) groups. We use 4/5ths for building the model, and 1/5th for evaluating. Then we move on to the next row and build another model. The average performance on the test data is an estimate of how a model might perform.
BTW, I mentioned "stratified" earlier. This means we maintain the proportions of positive and negative cases in the test and training sides of each row. Without it, all the positive cases could end up in the training set, and you've got nothing to test against.
To figure out the hyperparameters, we follow the same pattern on each of the splits, which is the smaller grid of yellow and blue boxes (an inner split).
We train models with different settings using that smaller training data set, and pick the best performing based on the inner test sets. Using those "best" hyperparameters, we train on all the training data from the original "outer" split, and evaluate it on the (green) test data from the outer split.
And so on down all the 5 (k) splits. Pretty involved. The result is getting a realistic estimate of how well a model would work for real, if we trained it using cross-validation on all the data.
* * *
If you like code, the scikit-learn project has a compact example.
